What’s in an image?
Embryonic stem cells are becoming something of a clinical holy grail – but is quantitative imaging an essential development for the quality control of human embryonic stem cells? We find out
Embryonic stem cells are becoming something of a clinical holy grail – but is quantitative imaging an essential development for the quality control of human embryonic stem cells? We find out
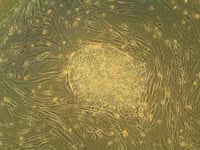
Qualitative examination of the cells is usually carried out by in situ staining and quantitative analysis carried out by flow cytometery. The images obtained from in situ staining give an indication of the state of one or two colonies, but this is not representative of the whole culture population. Flow cytometry on the other hand, looks at a sample of the whole population of cells, but not the individual colonies, nor does it give any information on the co-location of cells within a culture. Producing samples of hES cells for analysis on the flow cytometer can be problematic since the production of a single cell suspension results in up to 60% cell death (data not shown).
hES cells are commonly passaged using manual dissociation. Enzymatic treatment of these cells to obtain a single cell suspension results in large clumps of dead cells that need to be removed prior to use on the flow cytometer. The loss of a high number of cells from the analysis could lead to skewed results. Microscopy based fluorescent cytometry of fixed hES cell colonies can overcome these issues by allowing the automated processing of multi-labelled samples in situ. Automated image acquisition provides the capacity for thousands of images to be recorded. The software algorithms can identify the cells and the fluorescent pattern of the cells is then analysed, quantified and represented in dot plot format2. One system that performs this acquisition and analysis of images is the TissueFaxs (TissueGnostics, Austria).
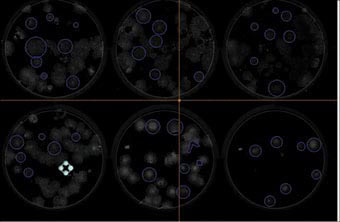
The TissueFaxs is a high quality fluorescent microscope with a modified automated stage that enables the acquisition of large numbers of images. The inverted microscope allows imaging of up to four slides or one multiwell plate at a time. A preview mode at x5 magnification (Figure 1) allows the plate to be imaged at low resolution and regions of interest can then be identified and imaged at higher resolution. The microscope is automated and will image a whole 6 well plate at x20 magnification on two fluorescent channels in 2 hours. Images can be stitched together to give high resolution images of an entire well. Figure 2 shows the stitched image created from over 3000 images at x20 magnification, of the whole well of a six well plate (approximately 9.2cm2). The system captures not only the morphology of the colonies, but also the location and number of the colonies present. The colour camera with the brightfield settings allows the acquisition of samples prepared for histology or non-fluorescent markers.
 |
| Figure 1: Stem cells grown on Matrigel, a x5 preview of DAPI to show the colonies of cells. The blue circles are the regions of interest to be acquired |
The software also measures other parameters such as the size of the nucleus and the number of cells/mm2, providing useful data for quality control. Figure 4 shows the data stored for each of the cells identified in the DAPI channel, similar data is stored for all the fluorescent channels imaged.
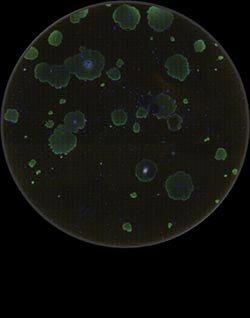 |
| Figure 2: Shef-7 cells grown on Matrigel. The stitched image is made of over 3000 x20 images. DAPI is used as the cell marker. Green fluorescence shows the cells positive for the marker Tra-1-60 found on hES cells |
The main benefits of “back-gating” or “forward-linking” are that data analysed is known to arise from cells that have been correctly identified, the negative cut-off is accurate and, any unexpected data can be traced back to the image it originated from for clarification.
To compare the data obtained from flow cytometry and the TissueFaxs, one plate of cells was harvested for flow cytometry and one was fixed for in situ staining. The cells were then stained and analysed. Figure 7a shows that whilst the data show similarities the difference between the TissueFaxs and the flow cytometer in terms of positive cells can be around 20%. In stem cell cultures substantial variation is observed in the number of cells per colony and the proportion of differentiated cells present. A 20% difference could simply reflect plate to plate variation rather than the sensitivity of either system. The percentage of differentiated cells indicated by SSEA1 expression was higher in the flow cytometry sample. This is possibly due to the areas imaged on the TissueFaxs being the colonies (the cells of primary interest) and SSEA1 expressing cells are more likely to be found on the periphery of the colonies and as single cells between the colonies.
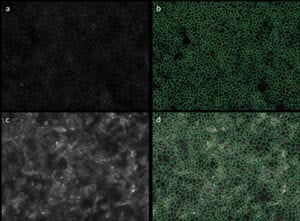 |
| Figure 3: a, DAPI staining of the nucleus. b, Identification of the nucleus by the software. c, FITC labelled stem cell marker (tra-1-81). d, area where the fluorescent signal is measure for each cell |
Based on our preliminary data, the sensitivity of the TissueFaxs may not be equivalent to flow cytometry. However, in our hands the TissueFaxs data appears to be potentially more robust and therefore more reliable for use in cell culture quality control. Comparison of in situ staining with the data from flow cytometry is not directly comparable as a large number of cells may be lost from the flow cytometry analysis. One disadvantage of the TissueFaxs is that data from cells in thick multicellular aggregates have to be removed from the TissueFaxs analysis. Nonetheless, our preliminary data indicates that in situ staining can provide valuable additional data not obtained from flow cytometry. In particular the variation in the number of cells from well to well, the density of cells/mm2, and most importantly the morphology of the colonies is recorded. In our view this extra data could prove indispensable where accurate quality control data is required for hES cell cultures. Changes in the density and morphology of cells may have a considerable affect on downstream assays and procedures, and we propose that simply qualitative evaluation of the expression of the stem cell markers is not sufficient to establish that the cells used as the starting point for different experimental work were equivalent.
 |
| Figure 4: Example images with the cells identified by the software. Highlighting an event brings up the event data for the fluorescent channel |
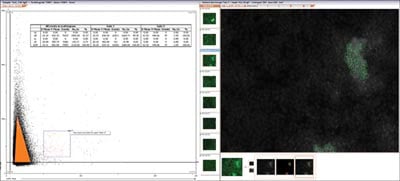 |
Figure 5: The dot plot of the events from one well stained with Tra-1-60. Gate 1 in orange shows single cells. Gate 2 in red has been 'back-gated' i.e. all the events in Gate 2 are identified in the images they were created from |
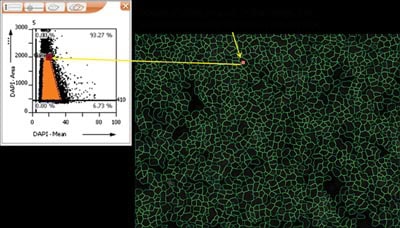 |
Figure 6: The DAPI image of the cells with the nucleus identified, with the linked data to the dotplot |
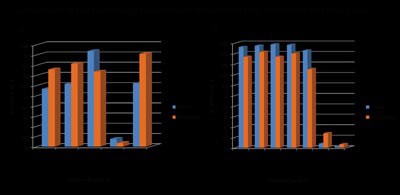 |
Figure 7: The percentage of positive cells detected with flow cytometry and the TissueFaxs. a, Comparison of cells stained in situ on the TissueFaxs and cells processed to single cells for flow cytometry (Guava) analysis. b, Single cell suspension stained for flow cytometry analysis analysed on the TissueFaxs and flow cytometer |




